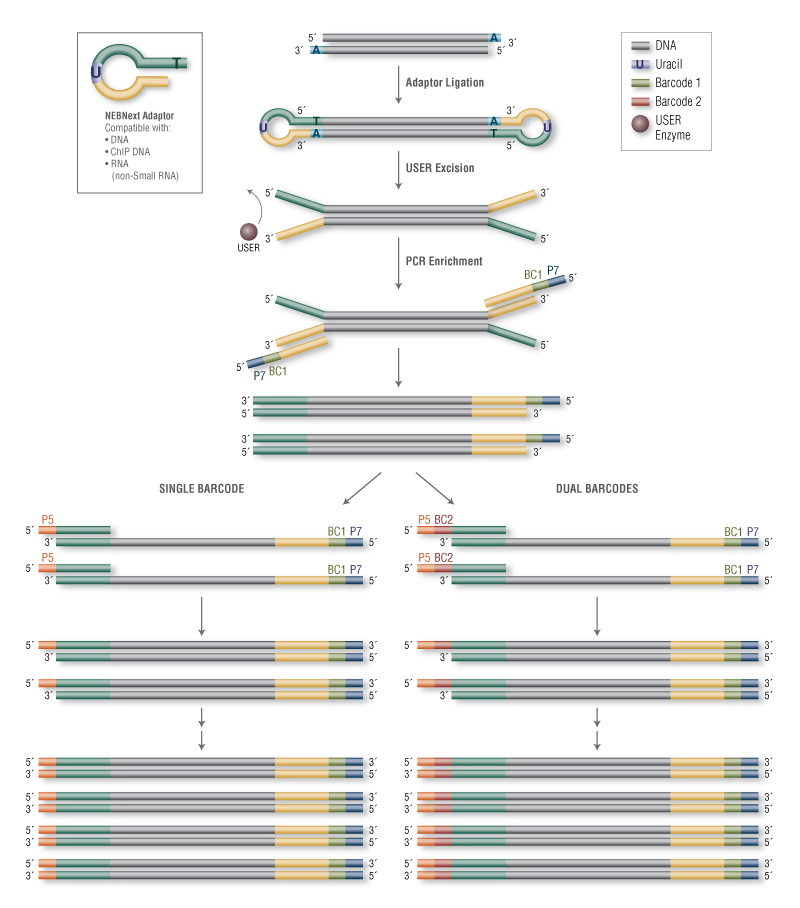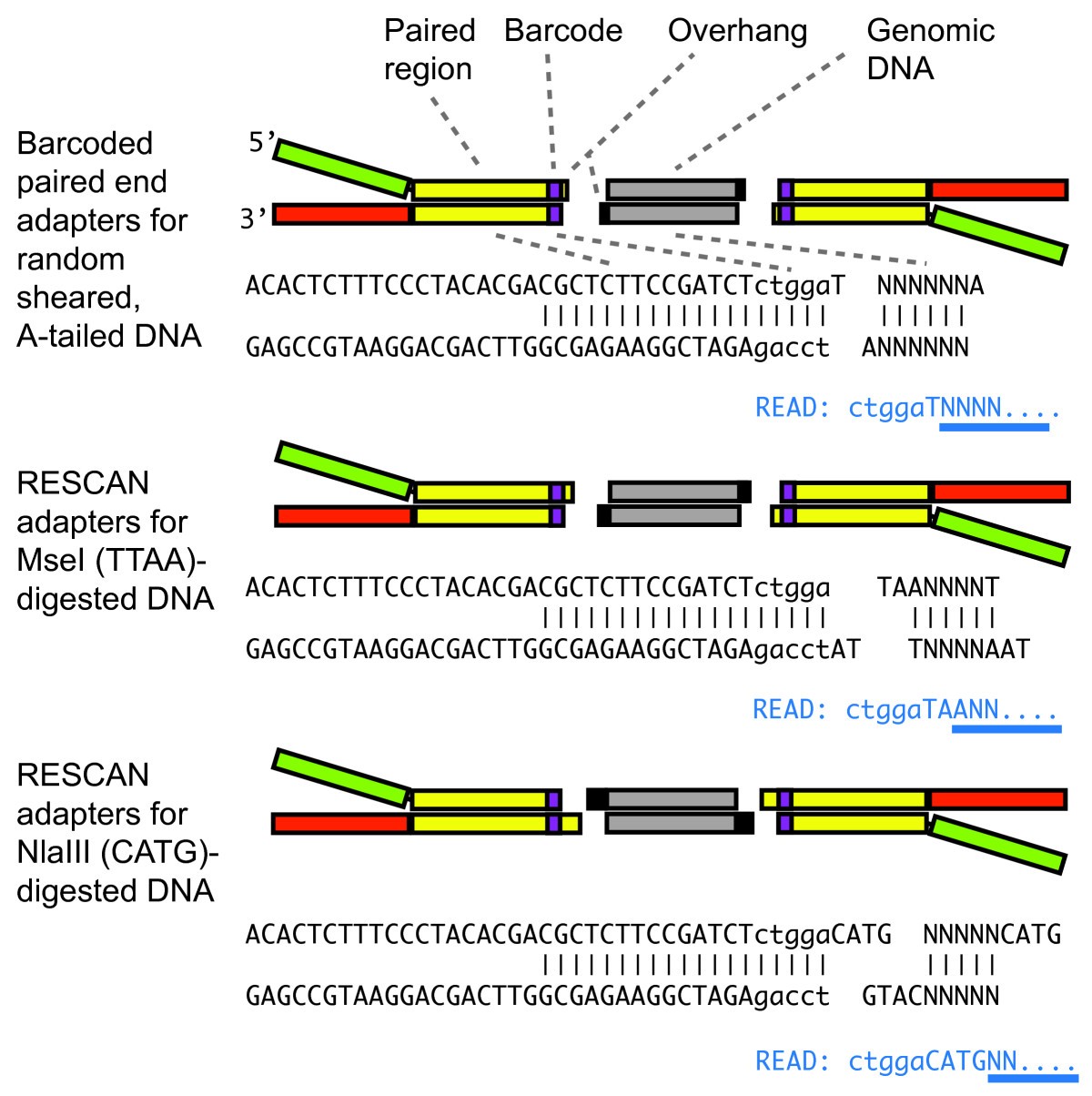
Reference genome-independent assessment of mutation density using restriction enzyme-phased sequencing | BMC Genomics | Full Text

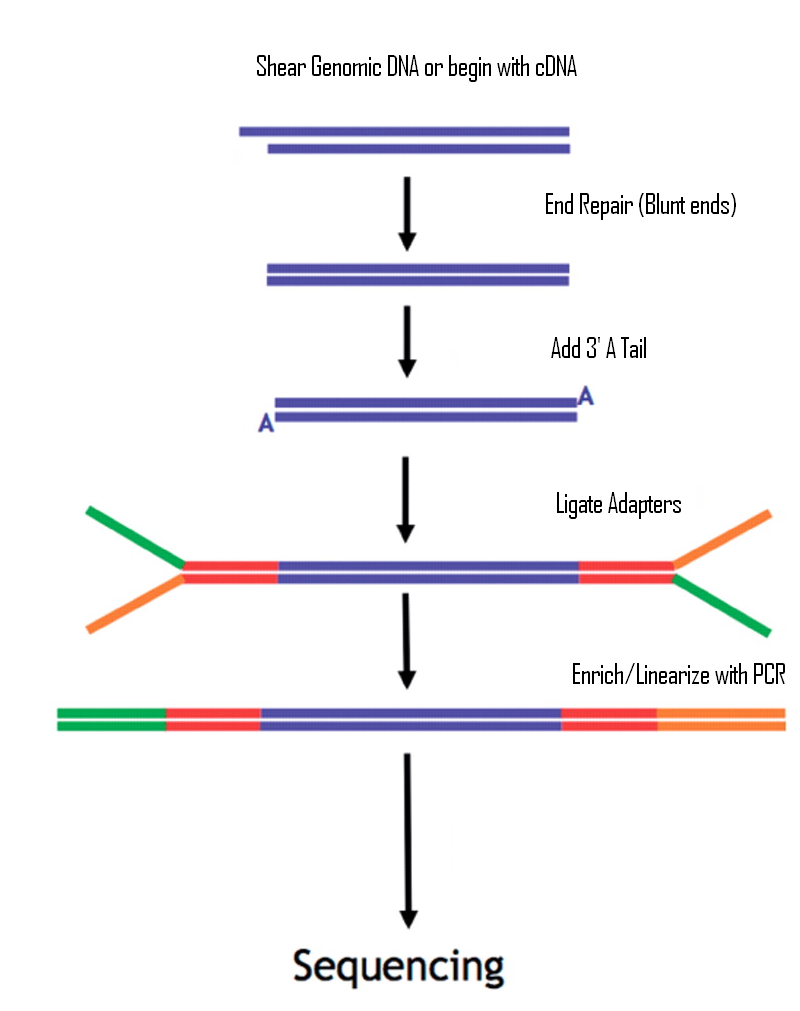
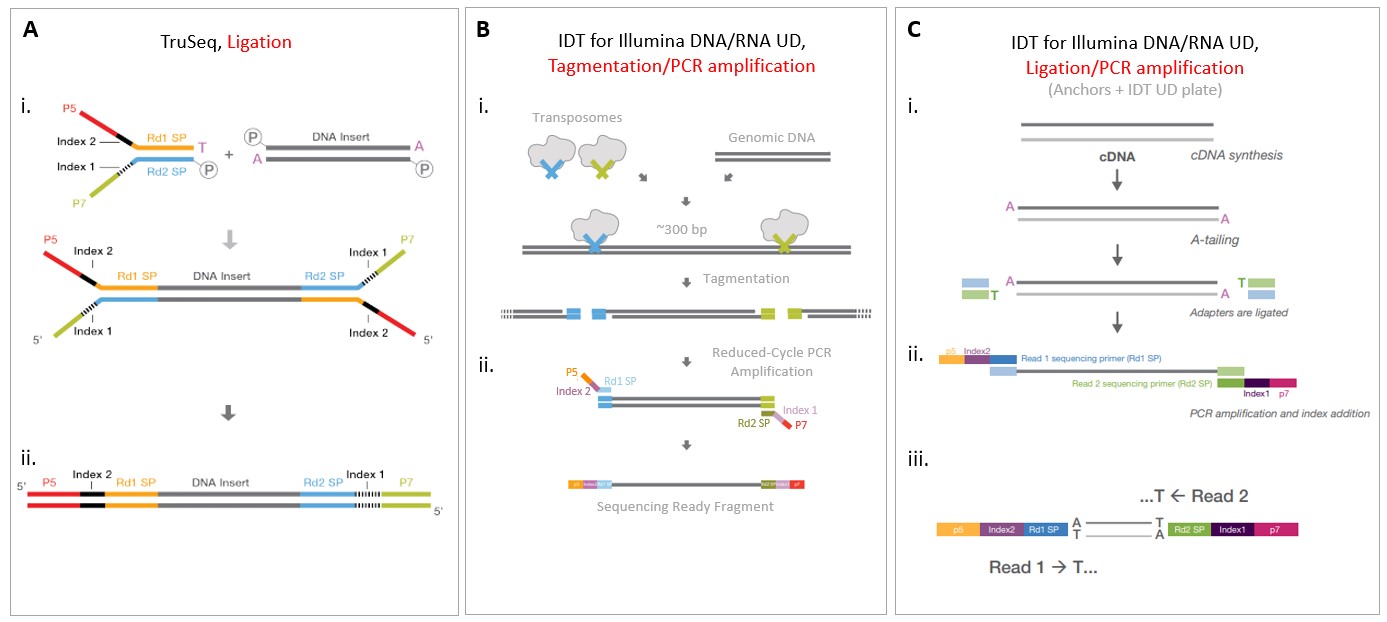
Preparation of DNA Sequencing Libraries for Illumina Systems—6 Key Steps in the Workflow | Thermo Fisher Scientific - US
Illumina Sequencing Sample Preparation for use with CRISPRi/a-v2 Libraries Overview This protocol describes the four steps requi

Preparation of Amplicon Libraries for Metabarcoding of Marine Eukaryotes Using Illumina MiSeq: The Adapter Ligation Method | SpringerLink
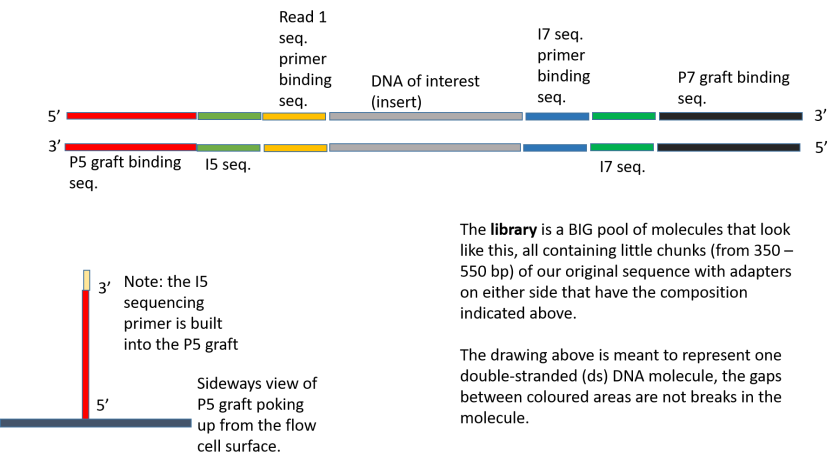
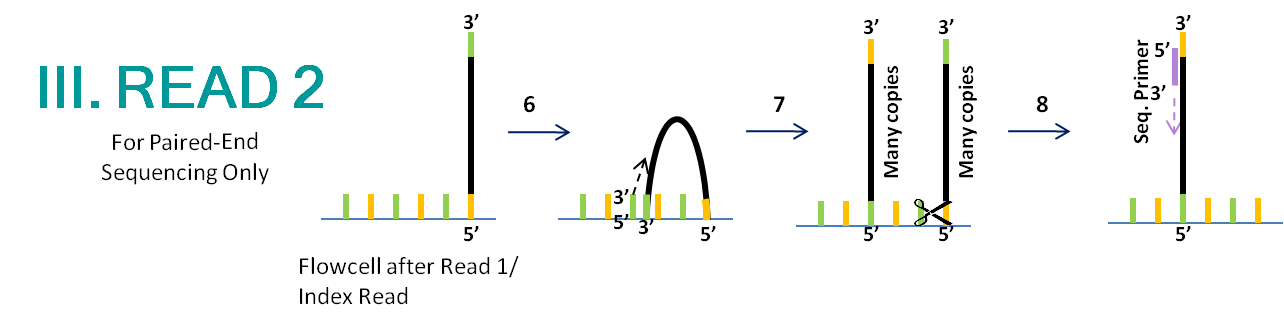
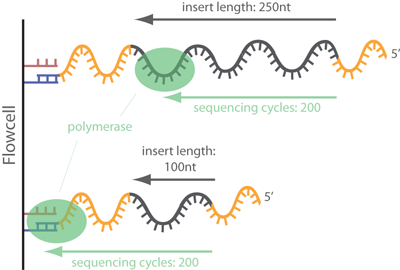
Illumina Sequencing (for Dummies) -An overview on how our samples are sequenced. – kscbioinformatics

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems

Overview of the different methods used for adapter addition to antibody... | Download Scientific Diagram

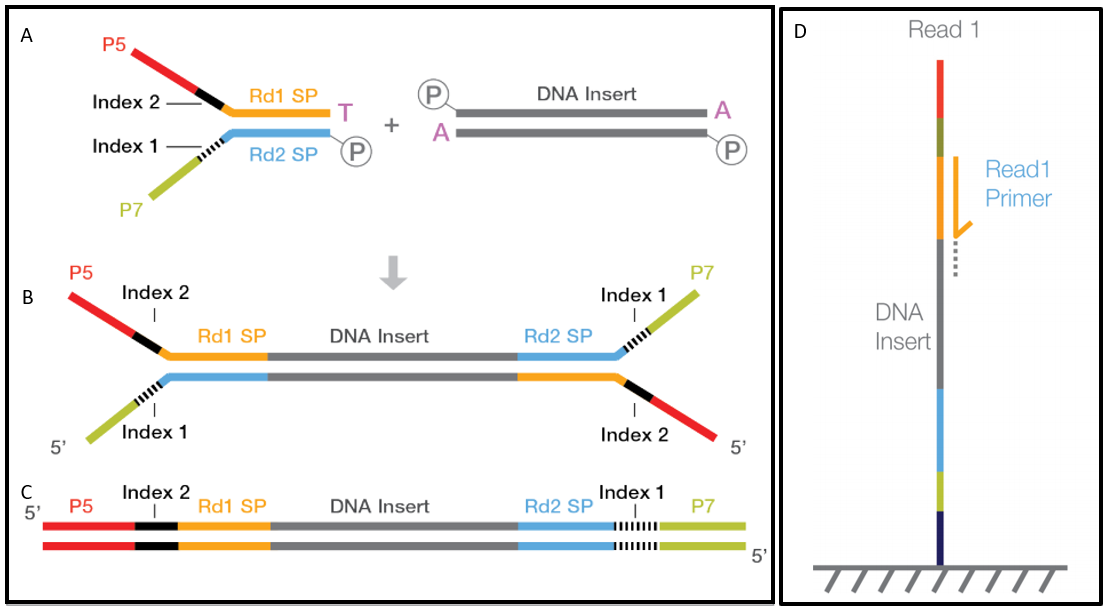


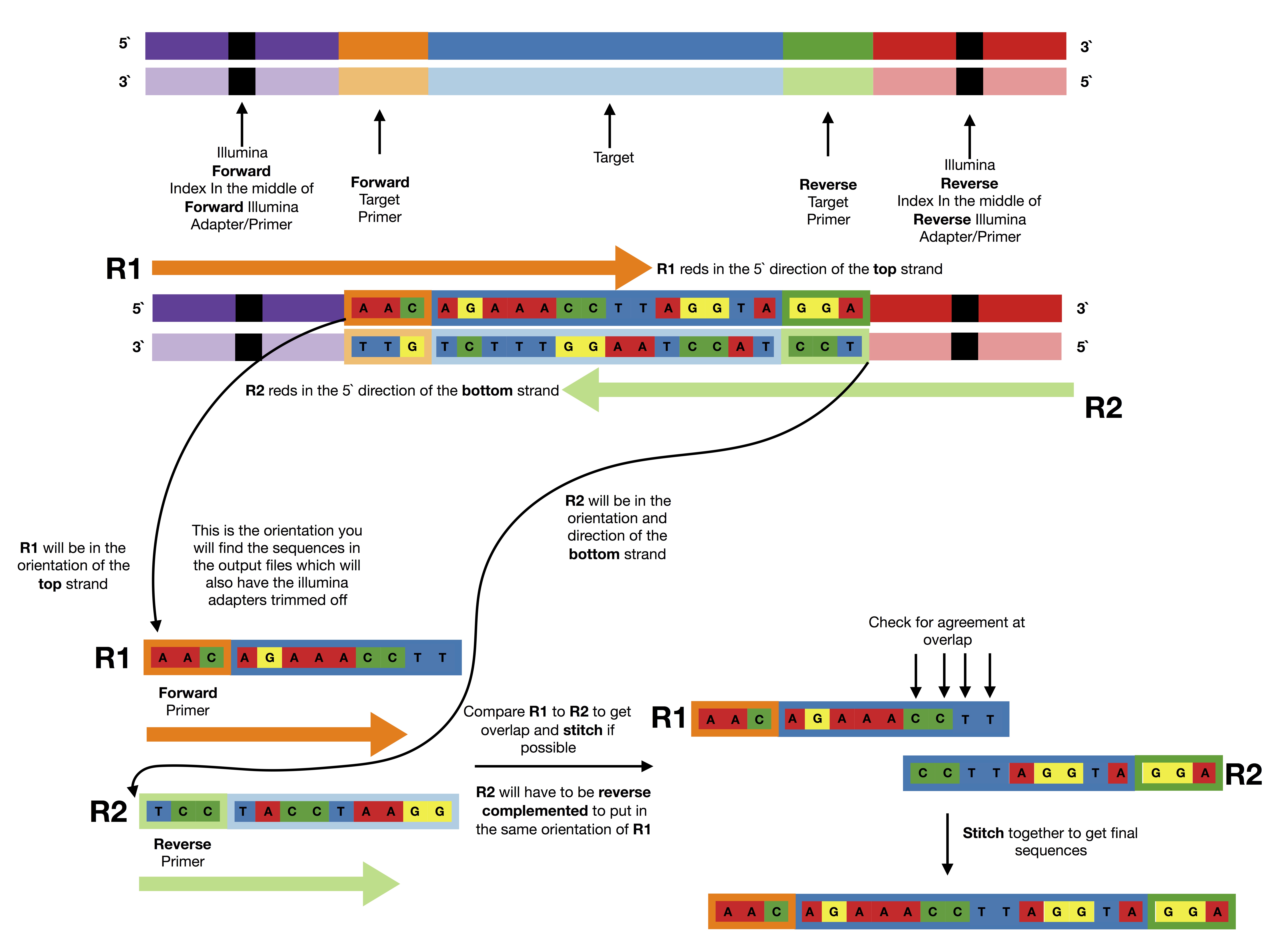
![PDF] leeHom: adaptor trimming and merging for Illumina sequencing reads | Semantic Scholar PDF] leeHom: adaptor trimming and merging for Illumina sequencing reads | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/247973613dfa3c42eb437ec9f5269202595f96af/2-Figure1-1.png)