![Comparative plastomics of Amaryllidaceae: inverted repeat expansion and the degradation of the ndh genes in Strumaria truncata Jacq. [PeerJ] Comparative plastomics of Amaryllidaceae: inverted repeat expansion and the degradation of the ndh genes in Strumaria truncata Jacq. [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/12400/1/fig-1-full.png)
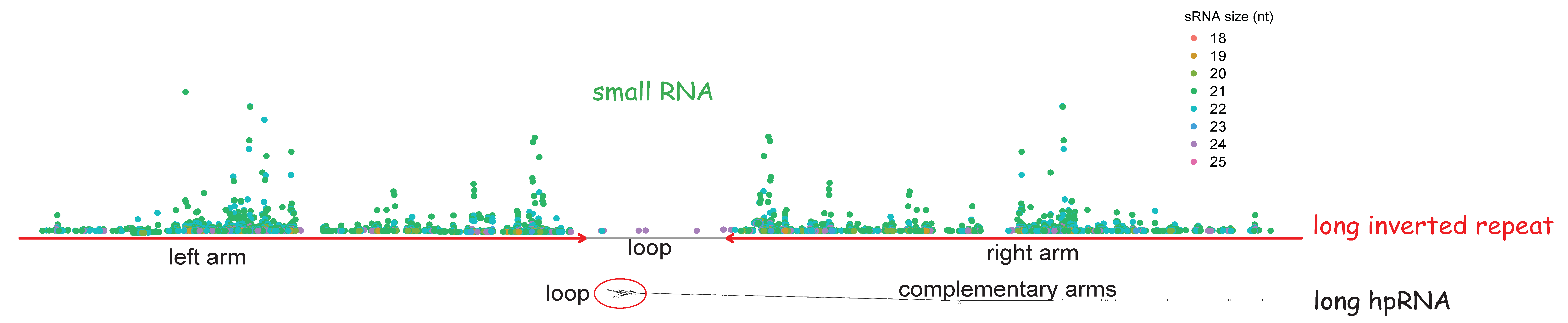
Comparative plastomics of Amaryllidaceae: inverted repeat expansion and the degradation of the ndh genes in Strumaria truncata Jacq. [PeerJ]
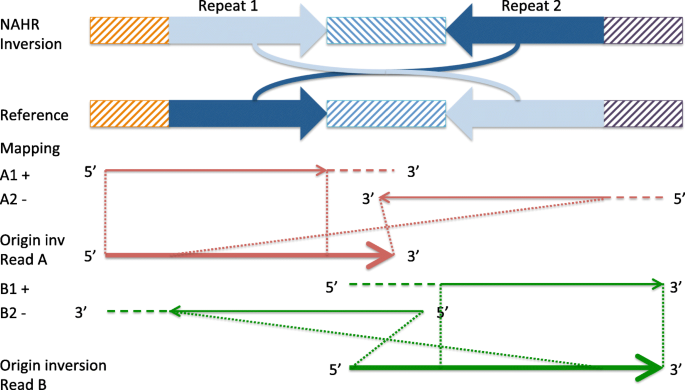
npInv: accurate detection and genotyping of inversions using long read sub-alignment | BMC Bioinformatics | Full Text

Large inverted repeats identified by intra-specific comparison of mitochondrial genomes provide insights into the evolution of Agrocybe aegerita - ScienceDirect
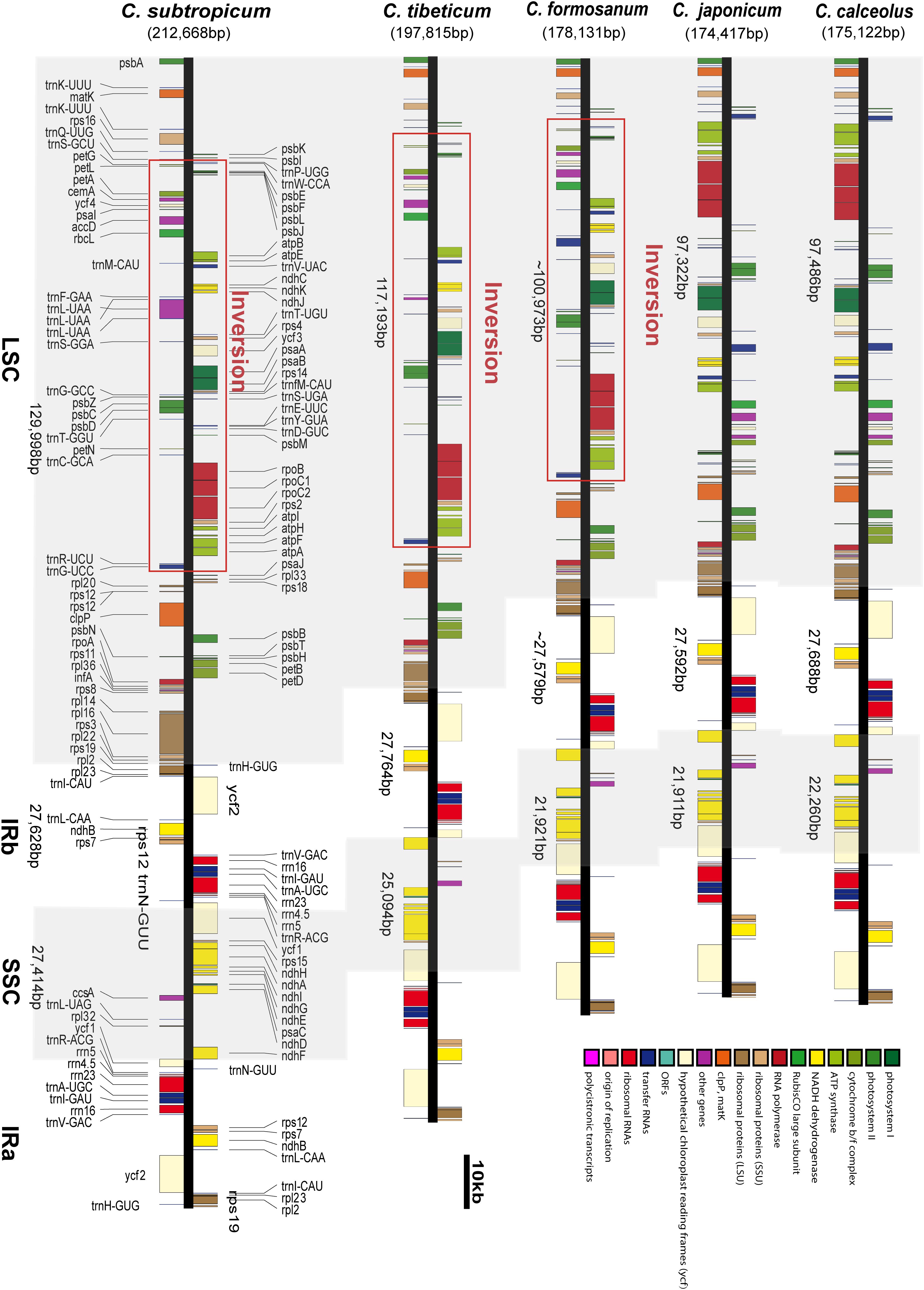
Frontiers | Chloroplast Genomes of Two Species of Cypripedium: Expanded Genome Size and Proliferation of AT-Biased Repeat Sequences

Correlations between long inverted repeat (LIR) features, deletion size and distance from breakpoint in human gross gene deletions | Scientific Reports
detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | PLOS ONE
![PDF] detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | Semantic Scholar PDF] detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a20f6bea6d62205d58f8bcd56fa3411c859aad47/3-Figure1-1.png)
PDF] detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | Semantic Scholar
detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation | PLOS ONE